Channel set invariance for neural networks
Devlopment of a neural network architecture invariant to different channel sets contained in EEG recordings
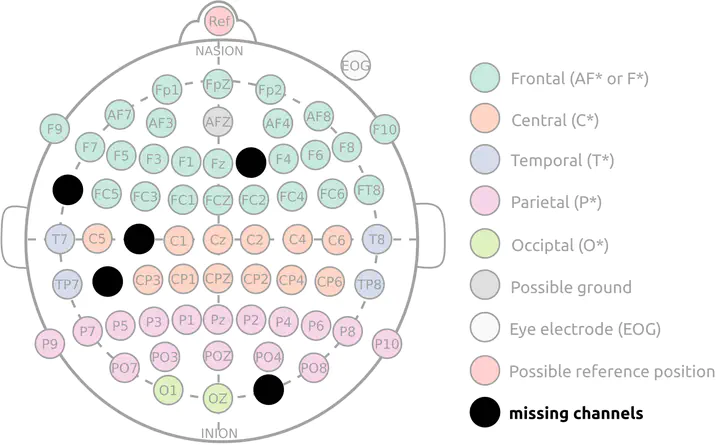
Problem
Despite the well-established standards for EEG electrode layout like the 10-10 montage, EEG recordings obtained in different labs or across studies are not straightforward to compare. There are usually important differences between the datasets, but also within them. These differences can be:
- the use of different channel sets or sensor placement routines
- different noise distributions, or changes of the noise structure over time
- temporarily unusable channels (electrical connection lost, faulty wire, …)
For this reason, spatial filters can hardly be re-used between sessions, let alone datasets, which is an obstacle for transfer learning approaches in EEG decoding problems. The existing solutions to do transfer learning with heterogeneous channel sets are:
- train a model on the common channel subset → restrictive, if a new recording has a faulty channel, as the whole model has to be re-trained without this channel
- interpolate the missing channels → obtained full channel set may not be full rank any more
- source reconstruction techniques → assumptions need to be made for source reconstruction, their choice is non-trivial
Objective
The objective of this project is to help in the development of a neural network architecture invariant to the set of channels used. Such an architecture would allow to realize an end-to-end learning approach across multiple datasets with heterogeneous channel sets. Ideally, this architecture would be able to:
- handle EEG signals containing an arbitrary number of channels
- produce results independently of the channels used
- reach the same performances as non-channel independent architectures (like EEGNet) when the channel set is fixed / complete
- seamlessly ignore corrupted channels, and degrade gracefully
- cope in real time with potential noise distribution shifts
Skills required
- Good programming skills in Python
- Experience with the pytorch library
- Optional: experience with deploying pipelines on GPU clusters
- Optional: experience with EEG/BCI data